
If you are interested in the average standard error of the difference between predicted means you can use “$avsed” from the generated object ’rcv.pv’. Hence, these values represent the expected mean yield performance of a given variety once it is ‘adjusted’ or ‘corrected’ by the other model terms, such as replicated in this case. In this instance, we have requested the adjusted means for all levels of variety, which are shown in the red rectangle together with their standard errors for the response variable means.
#ASREML R TRIAL#
R e 2 I n rI n c 2 r r c c ARMS Fusion 2007 Spatial Data Analysis Modelling of the autoregressive process RAD195 trial Dothistroma infection data (0-10 scores) surface plot. “Removing Spatial Variation from Wheat Yield Trials: A Comparison of Methods.” Crop Science, 86, pp. ASReml Workshop Harry Wu UPSC, Swedish University of Agriculture Science, Sweden CSIRO Plant Industry, Canberra, Australia.

#ASREML R SOFTWARE#
Stroup WW, Baenziger PS and Mulitze DK (1994). ASReml-R is powerful statistical analysis software specially designed for mixed models using Residual Maximum Likelihood (REML) to estimate the parameters in an R environment. ASReml-R is statistical software (and library in R) that fits linear mixed models (LMM) using REML methodology and calculates BLUE and BLUP values. Let’s see an example using the nin89 dataset. In this course we will focus on the fundamentals of using ASReml-R version 4 for the analyses of experimental data to fit models for biological studies. For predict.asreml(), your model term of interest will be referenced in the classify set. The output from ASReml-R forms predicted values for a factor and considers for the remaining variables, either user specified values of the remaining variables or average of these values. These predictions are sometimes called least-square means (LSMeans), but this term applies only to predictions from models without random effects. Genomic Best Linear Unbiased Prediction, or GBLUP, is a genomic selection method that uses genetic relations. Predictions are formed as an extra process after the final iteration and they are primarily used for generating tables of adjusted means for all levels of a given model factor. To see downloadable files click SHOW MORE below.
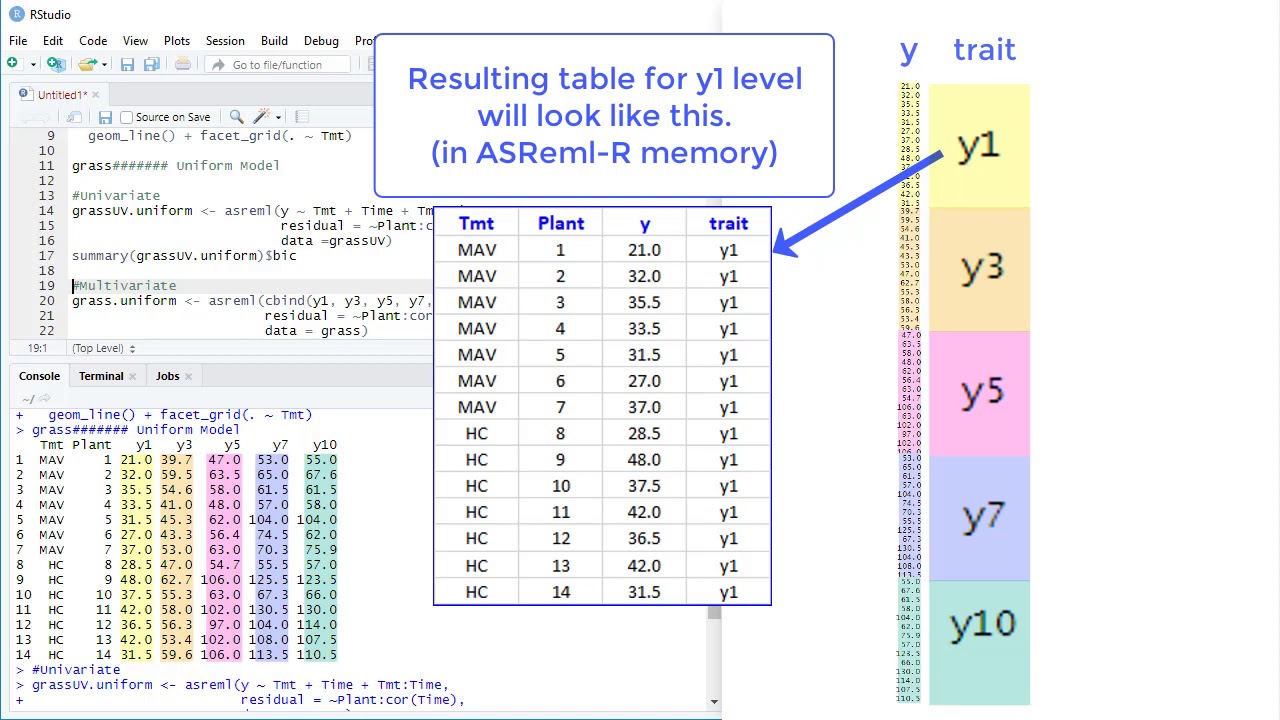
The “ predict.asreml ()” command in ASReml-R forms a linear function of the vector of fixed and random effects to obtain a predicted value for a factor of interest. I am using ASREML-R to fit unstructured (UN) and factor analytic (FA) model to explore complex structure of genotype by environment interaction in multienvironment yield data. You can find further information about this here. A “predict.asreml () ” function in ASReml-R ASReml-R 4 uses a new licensing system with different requirements from previous versions.


